Thresholding Image Stacks
2025-09-25
Source:vignettes/thresholding-image-stacks.Rmd
thresholding-image-stacks.RmdStacks of Images
50.tif is a TIFF file which is a stack of 50 images of a
bit of a cell taken over a short space of time (i.e. it’s a video).
img <- ijtiff::read_tif(system.file("extdata", "50.tif",
package = "autothresholdr"
))
#> Reading image from /home/runner/work/_temp/Library/autothresholdr/extdata/50.tif
#> Reading an 8-bit, float image with dimensions 100x100x1x50 (y,x,channel,frame) . . .
dim(img)
#> [1] 100 100 1 50Let’s display the first 3 frames:
and the last 3 frames:
You’ll notice that these images are almost identical. That’s because
they’re images of the same area taken very quickly one after another.
There are two ways to threshold an image stack like this. One is the
naiive way: find a threshold based on every pixel in the stack, then
pixels below the threshold are excluded (set to NA). Let’s
try that:
library(autothresholdr)
naiively_threshed_img <- apply_mask(img, "tri")
attr(naiively_threshed_img, "thresh") # The threshold chosen by "Triangle" is 4
#> [1] 4
#> attr(,"ignore_black")
#> [1] FALSE
#> attr(,"ignore_white")
#> [1] FALSE
#> attr(,"ignore_na")
#> [1] FALSE
#> attr(,"autothresh_method")
#> [1] "Triangle"
#> attr(,"class")
#> [1] "th" "integer"Now let’s display the first 3 frames and the last 3 frames:


If you look closely, you can see that the threshold mask is (slightly) different for different frames. Let’s highlight the pixels which are sometimes thresholded away, sometimes not:

So (unsurprisingly), it seems that around the edges of the cell (where the signal from the cell is more feint), the pixels are sometimes thresholded away, sometimes not. There are also some seemingly random pixels within the cell which are sometimes thresholded away, sometimes not.
Now, given that you know that the cell is more or less stationary and
you want the threshold to get rid of the non-cell bits and keep
the cell bits, its reasonable to assert that the mask should be
the same for every frame. It’s possible to apply the same mask to every
frame, and to compute this mask, it makes sense to incorporate
information from all of the frames. This is what
mean_stack_thresh() and med_stack_thresh() do.
mean_stack_thresh() computes the mask based on the mean of
all of the frames (gotten by calculating the mean intensity of the stack
at each pixel position). med_stack_thresh() uses the median
instead of the mean. They’re both very similar. If you don’t know which
one to use, just use mean_stack_thresh().
Let’s visualize them both:
ijtiff::display(mean_stack_thresh(img, "tri"))
ijtiff::display(med_stack_thresh(img, "tri"))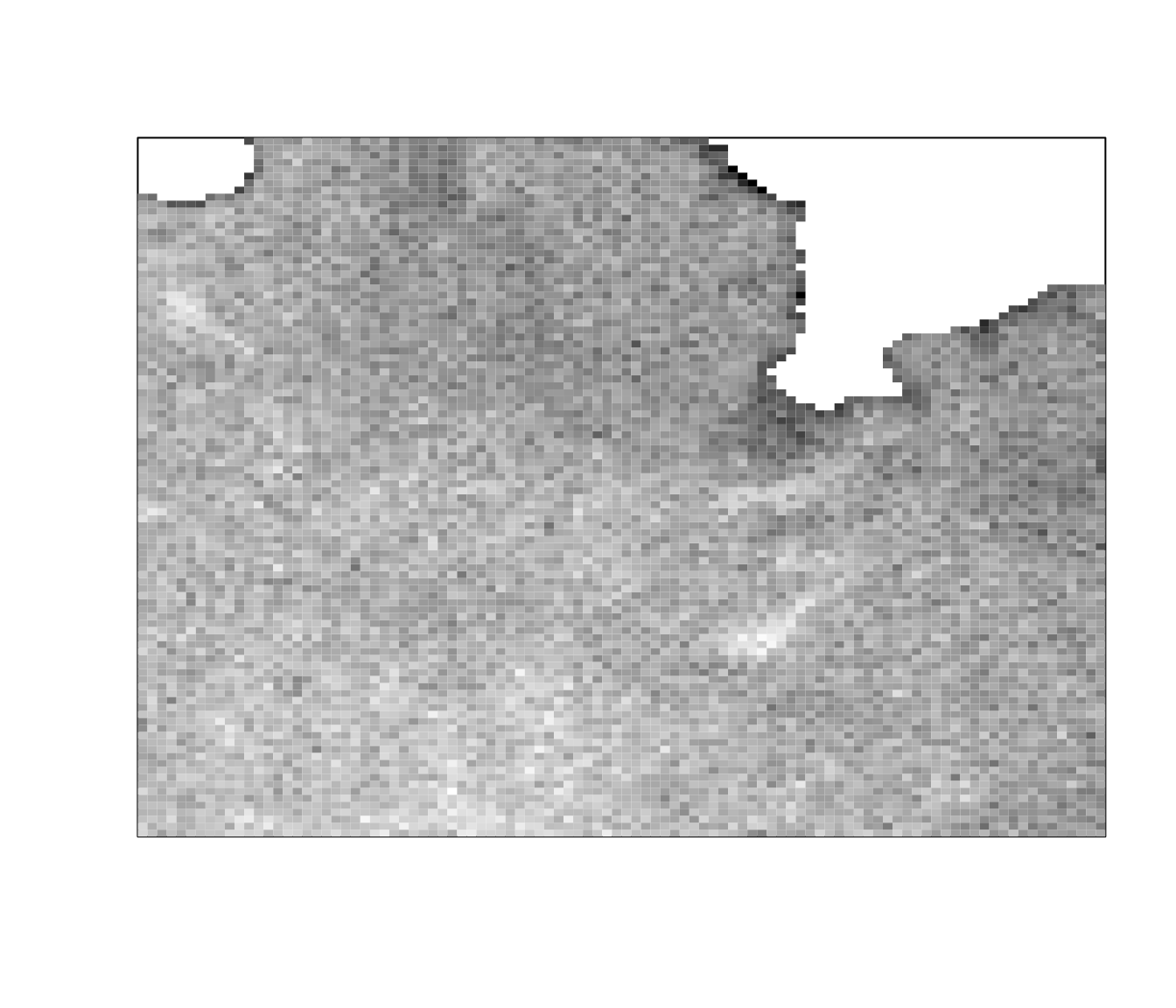
You can see that the results of mean_stack_thresh() and
med_stack_thresh() are similar but not identical. Both do a
fine job.
Note that if the cell (or whatever is recorded over the course of
several frames) is not stationary (or almost stationary), then
mean_stack_thresh() and med_stack_thresh() are
not appropriate.